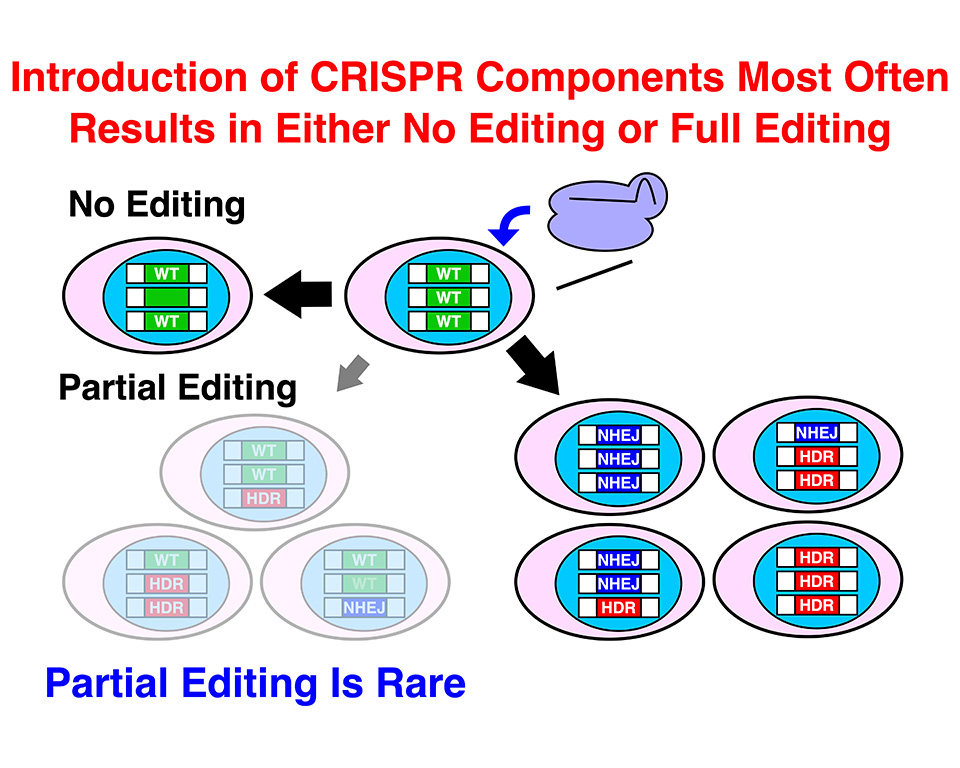
Genome editing has a great potential as a therapeutic approach. The most versatile genome editing tool, CRISPR-Cas9, can induce precise editing by inducing homology-directed repair (HDR), in which genome DNA is recombined with donor DNA. HDR precisely incorporates the donor DNA sequence into the genome. However, CRISPR-Cas9 also induces non-homologous end-joining (NHEJ) that generates random insertions and deletions with various sizes as by-products. It is important to grasp the frequencies and combinations of these different genome editing outcomes in single cells especially for therapeutic purposes. However, it has been difficult to analyze genome editing outcomes in individual cells. The current strategies to analyze genome editing results are mostly focused on cell populations but not single cells.
Gou Takahashi (staff scientist), Daiki Kondo (graduate student), Minato Maeda (graduate student), and Yuichiro Miyaoka (project leader) from the Regenerative Medicine project in collaboration with On-chip Biotechnologies Co., Ltd. established an efficient method to isolate human cell clones by using Single Particle isolation System (SPiS). SPiS is an automatic single cell dispenser with image analysis to precisely dispense one cell at a time. They isolated more than 2,600 clones from genome edited human cultured cell populations and analyzed their genome DNA sequences. It turned out that cells tend to undergo either no genome editing at all or full editing in all the target alleles in single cells (binary manner). They also found that in these fully edited cells, HDR tends to be accompanied by NHEJ. These findings are valuable information for genome editing therapies as well as basic life science.