
The Cdc48 ATPase (also known as p97) and its cofactor Ufd1-Npl4 captures and unfolds many polyubiquitylated proteins for proteasomal degradation, but how the Cdc48-Ufd1-Npl4 recognizes ubiquitin chains remained unclear. A collaborative research group, Dr. Shuya Fukai lab at the University of Tokyo and Dr. Yasushi Saeki lab, solved crystal structures of Npl4 complexed with Ufd1 and ubiquitin chains. The group identified a novel domain in Npl4 that specifically recognizes Lys48-linked ubiquitin chains, a primary signal for proteasomal degradation. The newly identified Npl4’s ubiquitin-binding site is needed for the interaction with polyubiquitylated proteins and Cdc48 activation, indicating the initial ubiquitin-binding site within the Cdc48-Ufd1-Npl4 complex. The group also determined the interaction surface of the Ufd1-Npl4 heterodimer, in which a flexible loop of Ufd1 is inserted into a hydrophobic groove of Npl4. Importantly, abolishing the Ufd1-Npl4 interaction destabilizes the Cdc48 complex and attenuates cell growth. Because the p97/Cdc48-Ufd1-Npl4 complex plays crucial roles upstream of the proteasome and has emerged as a promising target for developing anticancer drugs, the present structures provide a useful platform for the rational design of p97 inhibitors.
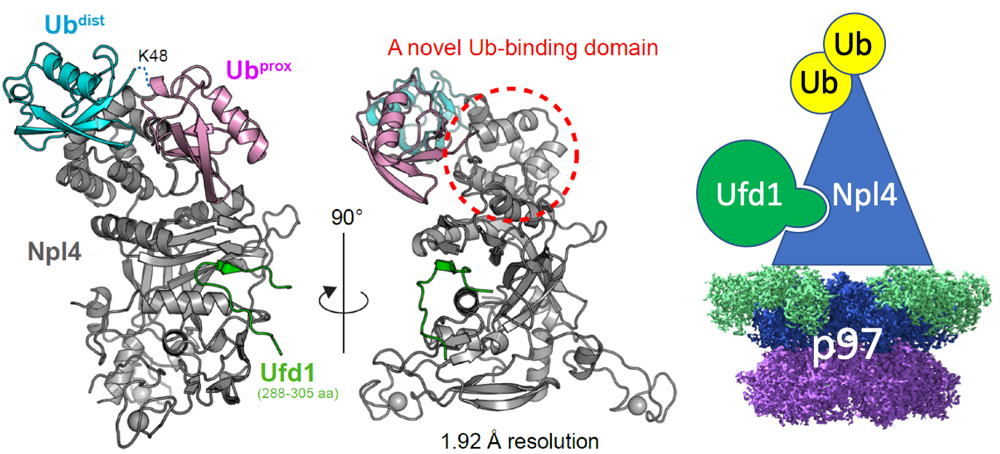
Left and middle: Crystal structure of Npl4 in complex with Lys48-linked diubiquitin and Ufd1. Right: Overall model of the Cdc48–Ufd-Npl4-diubiquitin complex. Distal and proximal ubiquitin molecules in ubiquitin chains are indicated as Ubdist and Ubprox, respectively.
Structural images are from the Protein Data Bank accession numbers 6JWI, 6JWI, and EMD-3295.