HOME > Topics2017 > 14 December 2017
14 December 2017
Naoko Niimi (Diabetic Neuropathy Project) published a paper on “A spontaneously immortalized Schwann cell line from aldose reductase-deficient mice as a useful tool for studying polyol pathway and aldehyde metabolism” in Journal of Neurochemistry.
Article publication on the establishment of a spontaneously immortalized Schwann cell line from aldose reductase-deficient mice
The increased glucose flux into the polyol pathway via aldose reductase (AR) in Schwann cells under high glucose conditions is recognized as a major contributing factor for the pathogenesis of diabetic neuropathy; however, little is known about the physiological roles of AR in the peripheral nervous system.
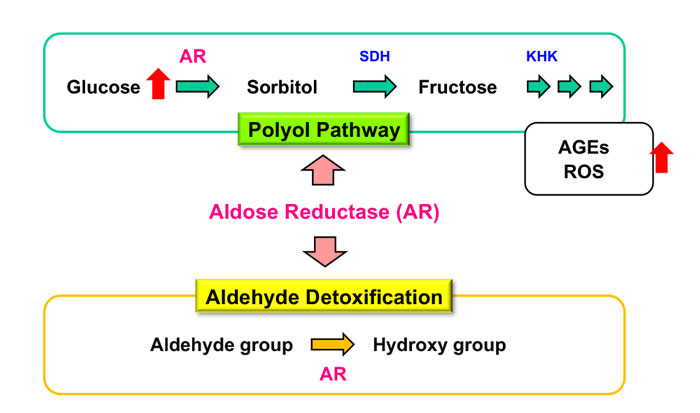
Aldose reductase (AR), the first enzyme in the polyol pathway, catalyzes the conversion of glucose to sorbitol. The escalation of the glucose flux into the polyol pathway via AR under diabetic conditions results in the increases of advanced glycation endproducts (AGEs) and reactive oxygen species (ROS), leading to the development of diabetic complications. On the other hand, AR catalyzes the reduction of reactive biogenic aldehydes and plays a role in aldehyde detoxification.
In collaboration with Hirosaki University, Sumitomo Dainippon Pharma Co., Ltd., Kyorin University and the University of Hong Kong, Dr. Naoko Niimi (Diabetic Neuropathy Project) and others of Tokyo Metropolitan Institute of Medical Science established spontaneously immortalized Schwann cell lines from long-term cultures of normal and AR-deficient C57BL/6 mouse peripheral nerves. These cell lines exhibited distinct Schwann cell phenotypes, such as spindle-shaped morphology, immunoreactivity for S100 protein, and synthesis and secretion of neurotrophic factors. Microarray and real-time RT-PCR analyses revealed significantly down-regulated mRNA expression of polyol pathway-related enzymes, sorbitol dehydrogenase and ketohexokinase, and significantly up-regulated mRNA expression of aldo-keto reductases and aldehyde dehydrogenases, in AR-deficient Schwann cells compared with those in normal cells. Exposure to reactive aldehydes (3-deoxyglucosone, methylglyoxal, and 4-hydroxynonenal) significantly up-regulated the mRNA expression of aldo-keto reductases AKR1B7 and AKR1B8 in AR-deficient Schwann cells, but not in normal cells. Because no significant differences in viability between these two cell lines after exposure to these aldehydes were detected, it can be assumed that the aldehyde detoxification is taken over by AKR1B7 and AKR1B8 in the absence of AR.
These cell lines will be useful tools for studying the physiological and pathological roles of AR.
- <Title of the paper>
- A spontaneously immortalized Schwann cell line from aldose reductase-deficient mice as a useful tool for studying polyol pathway and aldehyde metabolism
- <Authors>
- Naoko Niimi (Tokyo Metropolitan Institute of Medical Science)
Hideji Yako (Tokyo Metropolitan Institute of Medical Science)
Shizuka Takaku (Tokyo Metropolitan Institute of Medical Science)
Hiroshi Kato (Sumitomo Dainippon Pharma Co., Ltd.)
Takafumi Matsumoto (Sumitomo Dainippon Pharma Co., Ltd.)
Yasumasa Nishito (Tokyo Metropolitan Institute of Medical Science)
Kazuhiko Watabe (Kyorin University)
Kaori Ogasawara (Hirosaki University)
Hiroki Mizukami (Hirosaki University)
Soroku Yagihashi (Hirosaki University)
Sookja K. Chung (The University of Hong Kong)
Kazunori Sango (Tokyo Metropolitan Institute of Medical Science) - <Journal>
- Journal of Neurochemistry (December 2017)
http://onlinelibrary.wiley.com/doi/10.1111/jnc.14277/full

