Meet our scientists!
 Back
Back

Saki
Gotoh-
Saito
Saki
Gotoh-
Saito
Doctors have been using medicines to treat patients for thousands of years. However, different patients react differently to different medicines, reflecting the unique individuality of each patient. With the advent of whole-genome sequencing, it should theoretically become possible to map the differences between patients and tailor treatments to specific individuals. Saki Gotoh-Saito, a research scientist studying transcription at the Research Center for Genome and Medical Sciences, recently published a paper, Drug-induced cis-regulatory elements in human hepatocytes affect molecular phenotypes associated with adverse reactions, in Nature Communications. In this paper, Dr. Gotoh-Saito identifies variations in non-coding areas of the genome that affect our reactions to different medicines or drugs. Mutations in coding regions of our genomes (DNA sequences that are copied into mRNAs and translated into proteins) are well-known to produce altered proteins that affect different biological functions. However, mapping the effects of variations in regulatory regions (DNA regions that affect the amounts and types of mRNA transcripts made) on traits has been less common but is required for progress towards individualized medicine. We spoke to Dr. Gotoh-Saito about her work.
How did you first become interested in research and biology?
To tell you the truth, I didn’t like biology at all in High School, and I didn’t study it much. But right after graduating, I happened to read a book on the human genome that blew my mind! This book described how DNA is composed of just four bases that are formed into long chains that encode genes, and how these genes are transcribed into messenger RNAs, which are then translated into proteins that do all the work in cells, and how this is the basis of all life on earth. It opened up a whole new world for me that I hadn’t known existed! I was so amazed that I decided to major in Biology in college, and there, I realized that I really enjoy experiments.
Another major influence for me was studying abroad in the United States while I was in graduate school. The lab where I studied in Japan was very small, but during my time abroad, I had the opportunity to study nuclear receptors at a lab at NIH, which had great people and facilities. It was there that my interest in gene expression really deepened.
What do you enjoy most about research?
I enjoy the whole process! From thinking of a hypothesis that explains something, thinking of experiments to test that hypothesis, and obtaining results that tell me whether the hypothesis was correct or not. All of those steps are fascinating and fun!
How did you start your current work?
Dr. Kawaji, the head of the Research Center for Genome and Medical Sciences, allowed me to pursue any projects I wanted, so I decided to study the pregnane X receptor (PXR), which I’d studied when I was in the US. PXR is a ligand-activated transcription factor that belongs to the nuclear receptor superfamily. PXR has a flexible ligand-binding pocket that allows it to bind to a wide range of prescription drugs. Once these drugs binds to PXR, PXR becomes active and induces expression of target genes involved in drug metabolism and transport. One documented adverse effect of several drugs that activate PXR is vitamin D deficiency, but the mechanism by which these drugs cause this deficiency had been unknown. I thought that by studying the cis-regulatory sequences (DNA sites) where PXR binds and identifying their target genes, I could understand why different people metabolize drugs differently, and why some become deficient for vitamin D
Could you explain your paper in more detail?
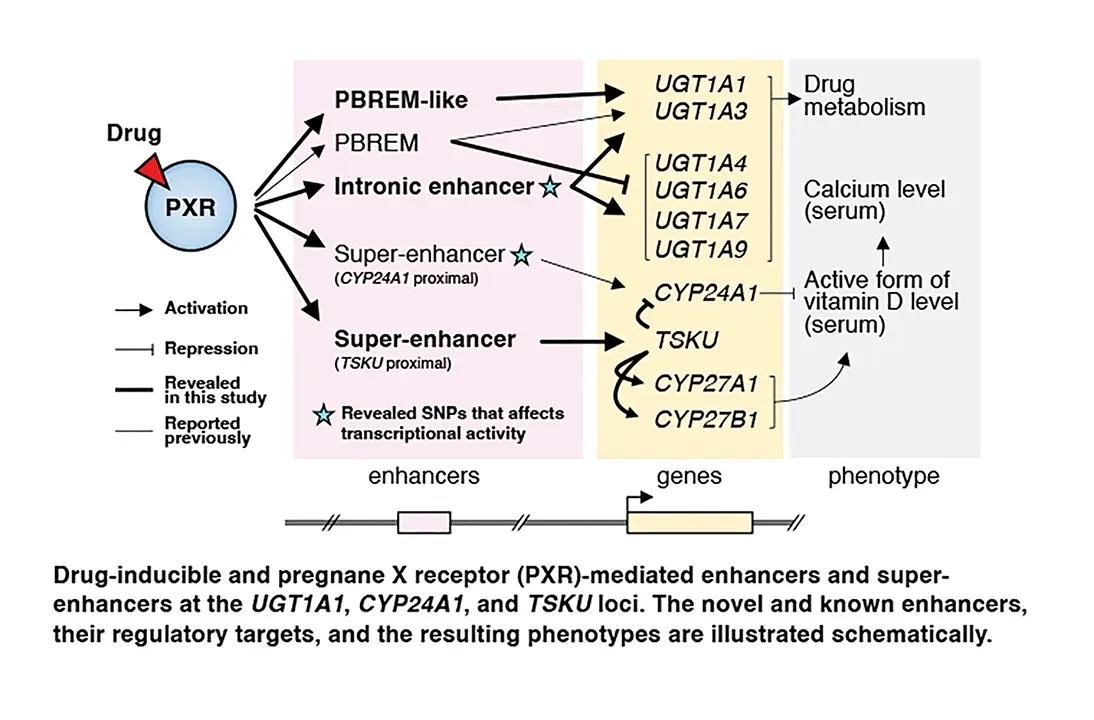
Sure. We used a method called cap analysis of gene expression (CAGE) to determine where transcription is activated in the genome when cells are treated with a drug, rifampicin. Rifampicin acts as a high affinity ligand for the human PXR. This activates PXR and causes it to bind to specific DNA sites called cis-regulatory elements (CREs). These CREs are located within enhancers that boost transcription of their target genes. When PXR binds to CREs, it increases expression of downstream genes to make messenger RNAs. They also drive expression of noncoding RNAs from enhancers called enhancer RNAs or eRNAs. Thus, by using CAGE to determine where transcription increases, we identified rifampicin-activated promoters, and we also identified thousands of enhancers where PXR is likely to bind.
We combined our results with two other types of data from various previous studies. The first was data from mapping studies that identified sites that PXR binds to. From this, we were able to identify CREs that bind to PXR and increase activity in the presence of rifampicin. The second was genome-wide association studies (GWAS), which show linkages between traits such as drug-induced vitamin D deficiency and single nucleotide polymorphisms (SNPs). Basically, GWAS studies allow us to map traits to specific regions of DNA. By combining these data, we were able to identify PXR enhancers that were likely associated with traits such as drug-induced vitamin D deficiency.
Finally, we were able to determine the function of our identified PXR enhancers by inhibiting or deleting them and measuring effects on their target genes.
Why is this work important?
People have different responses to drugs. For example, some people are susceptible to vitamin D deficiency when treated with certain drugs. A more serious case would be that some patients have an increased risk of neutropenia (an abnormally low number of neutrophils which increases the risk of serious infections) when treated with the anticancer drug irinotecan. We knew previously that these effects had something to do with PXR or genes associated with PXR, but not much more. Our work identifies the specific enhancers where PXR binds to regulate genes linked to a patient's susceptibility to neutropenia or vitamin D deficiency. Our work suggests that there are different variants of these enhancers that alter their binding affinity for PXR. This difference in binding directly affects gene expression, explaining why some people are more susceptible to severe side effects such as neutropenia or vitamin D deficiency than others. We haven’t proven this, but I think this idea makes sense and it helps us to understand why different medicines have different effects on different patients.
What are your plans for the future?
Right now, I am studying the function of a drug-inducible non-coding RNA. Messenger RNAs get translated into proteins, so everyone knows their importance, but non-coding RNAs including eRNAs don’t code for proteins so it’s still a mystery why they are made. I don’t think they are junk. They have a function and I’m currently studying one which binds to PXR. In any event, I really enjoy research so I’m planning on continuing to form hypotheses, conducting experiments, and obtaining results.
Interviewed by Jun Horiuch