Naoko Yoshizawa-Sugata and Hisao Masai at Genome Dynamics Project published an article entitled “Loss of full-length DNA replication regulator Rif1 in two-cell embryos is associated with zygotic transcriptional activation.” in the journal “Journal of Biological Chemistry”. The paper was released on line on November 1, 2021.

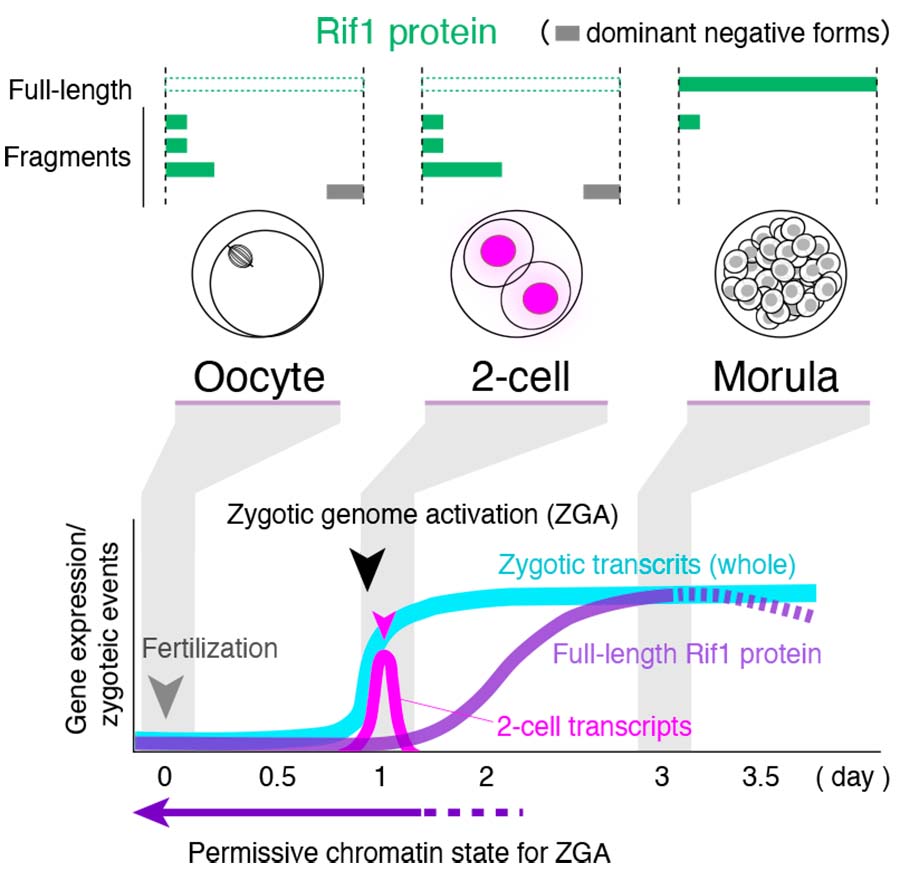
Rif1 regulates DNA replication timing and double-strand break repair, and its depletion induces transcriptional bursting of 2-cell (2C) zygote-specific genes in mouse ES cells. However, how Rif1 regulates zygotic transcription is unclear. We show here that Rif1 depletion promotes the formation of a unique Zscan4 enhancer structure harboring both histone H3 lysine 27 acetylation (H3K27ac) and moderate levels of silencing chromatin mark H3K9me3. Curiously, another enhancer mark H3K4me1 is missing whereas DNA methylation is still maintained in the structure, which spreads across gene bodies and neighboring regions within the Zscan4 gene cluster. We also found by function analyses of Rif1 domains in ES cells that ectopic expression of Rif1 lacking N-terminal domain results in upregulation of 2C transcripts. This appears to be caused by dominant negative inhibition of endogenous Rif1 protein localization at the nuclear periphery through formation of hetero-oligomers between the N- terminally truncated and endogenous forms. Strikingly, in murine 2-cell embryos, most of Rif1-derived polypeptides are expressed as truncated forms in soluble nuclear or cytosolic fraction, and are likely non-functional. Toward the morula stage, the full-length form of Rif1 gradually increased. Our results suggest that the absence of the functional full-length Rif1 due to its instability or alternative splicing and potential inactivation of Rif1 through dominant inhibition by N-terminally truncated Rif1 polypeptides, may be involved in 2C-specific transcription program. The dynamics of chromatin structure-regulatory proteins in early developmental process are not known precisely so far, and this study has elucidated one of the mechanisms that the loss of Rif1 during early stages of mouse preimplantation embryos may be related to zygotic transcriptional activation.