- HOME
- Cancer Immunology Project
Cancer Immunology Project
A comprehensive study of cancer immunity and its application to cancer treatment
Achievements in 2024
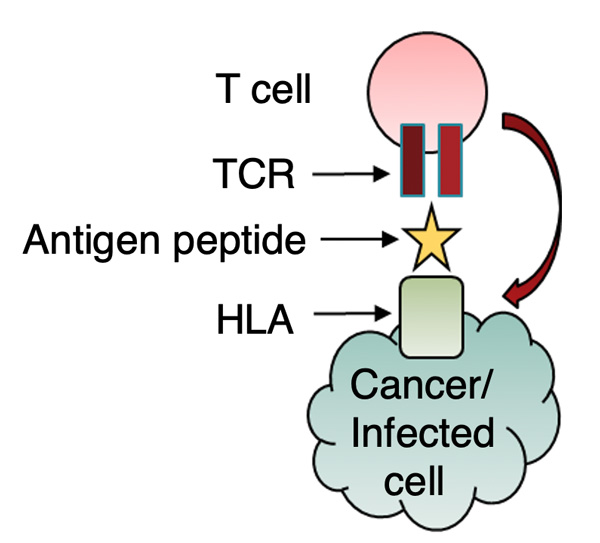
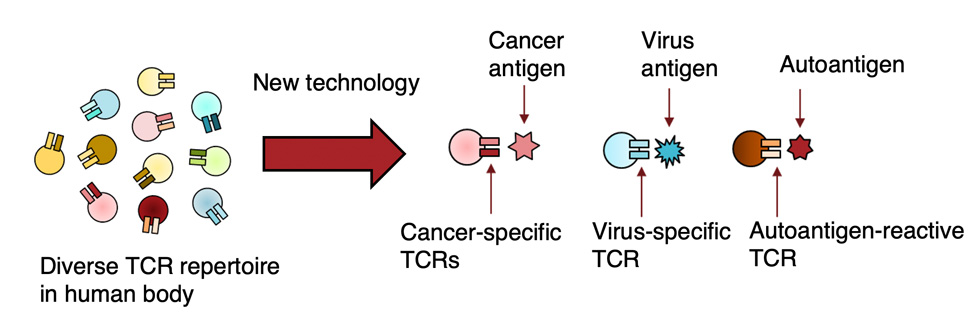
In previous work, we developed a high-throughput technology capable of determining TCR and antibody sequences at the single-cell level, which was published in PNAS (2020) and Science Advances (2020). However, this technology was unable to determine the antigen specificity of TCRs and antibodies. In 2024, we independently developed a proprietary platform that can identify antigen-specific TCRs as well as TCR-like antibodies. Using this technology, we are generating various cancer-specific TCRs and TCR-like antibodies. In parallel, we established cell lines derived from cancer patients to provide a clinically relevant model system. We are currently evaluating the cancer-killing efficacy of these TCRs and antibodies using both in vitro and in vivo models.
Publications
Key papers
- H Tanno et al. (2020) “A Facile Technology for the High Throughput Sequencing of the Paired VH:VL and TCRβ:TCRα Repertoires.” Science Advances. 6(17):eaay9093
- H Tanno et al. (2020) “Determinants governing T cell receptor α/ββ-chain pairing in repertoire formation of identical twins” PNAS. 117(1):532-540.

